Hello all,
I am implementing PCE in chaospy with Python and Uqlab with MATLAB, but I get different outcomes. In Python, the LOO is around 2%, but in MATLAB, it is 100%. I am wondering what is wrong with the UQLab MATLAB code? Here is my code:
MATLAB code:
clear
uqlab
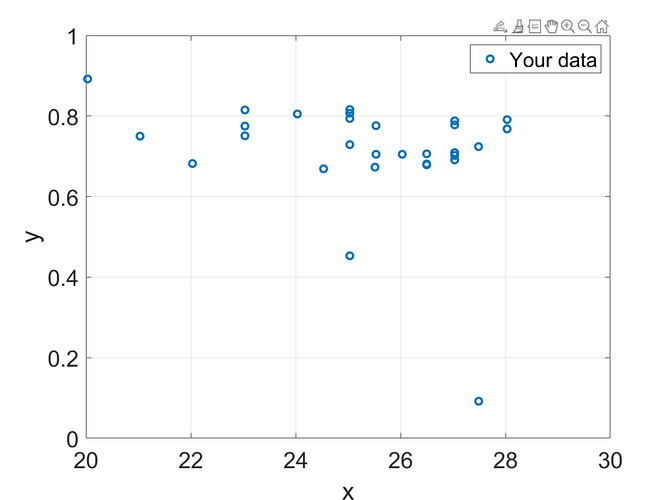
data = [
25.532, 0.776;
27.487, 0.724;
27.032, 0.788;
25.512, 0.673;
27.489, 0.092;
25.032, 0.453;
26.498, 0.706;
23.032, 0.775;
27.032, 0.702;
25.532, 0.705;
27.032, 0.691;
24.532, 0.669;
25.032, 0.794;
27.032, 0.709;
26.498, 0.681;
25.032, 0.807;
24.032, 0.805;
28.032, 0.791;
22.032, 0.682;
26.032, 0.705;
23.032, 0.751;
21.032, 0.750;
25.032, 0.816;
23.032, 0.815;
26.498, 0.679;
25.032, 0.729;
20.032, 0.892;
27.032, 0.778;
28.032, 0.768
];
% Use only the first and last columns
x = data(:, 1);
y = data(:, 2);
% Define the input marginals
InputOpts.Marginals(1).Name = 'X1';
InputOpts.Marginals(1).Type = 'Gaussian';
InputOpts.Marginals(1).Parameters = [mean(x), std(x)];
myInput = uq_createInput(InputOpts);
% Define the PCE metamodel
MetaOpts.Type = 'Metamodel';
MetaOpts.MetaType = 'PCE';
MetaOpts.ExpDesign.X = x;
MetaOpts.ExpDesign.Y = y;
MetaOpts.Degree = 2; % Ensure the same polynomial degree
MetaOpts.TruncOptions.MaxInteraction = 1; % Ensure the same truncation options
myPCE = uq_createModel(MetaOpts);
uq_print(myPCE);
##########################
Python code:
import numpy as np
import chaospy as cp
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import LeaveOneOut
from sklearn.metrics import mean_squared_error
# Provided data
data = np.array([
[25.532, 0.776],
[27.487, 0.724],
[27.032, 0.788],
[25.512, 0.673],
[27.489, 0.092],
[25.032, 0.453],
[26.498, 0.706],
[23.032, 0.775],
[27.032, 0.702],
[25.532, 0.705],
[27.032, 0.691],
[24.532, 0.669],
[25.032, 0.794],
[27.032, 0.709],
[26.498, 0.681],
[25.032, 0.807],
[24.032, 0.805],
[28.032, 0.791],
[22.032, 0.682],
[26.032, 0.705],
[23.032, 0.751],
[21.032, 0.750],
[25.032, 0.816],
[23.032, 0.815],
[26.498, 0.679],
[25.032, 0.729],
[20.032, 0.892],
[27.032, 0.778],
[28.032, 0.768]
])
# Split input and output data
X = data[:, 0].reshape(-1, 1)
y = data[:, 1]
# Define the distribution of the input variable (assuming normal distribution)
mean = np.mean(X)
std = np.std(X)
distribution = cp.Normal(mean, std)
# Create a joint distribution
joint_dist = cp.J(distribution)
# Define the polynomial expansion
polynomial_expansion = cp.generate_expansion(2, joint_dist)
# Evaluate the polynomial expansion at the data points
X_poly = cp.call(polynomial_expansion, X.T).T
# Initialize Leave-One-Out cross-validator
loo = LeaveOneOut()
# Initialize lists to store results
y_true = []
y_pred = []
# Perform Leave-One-Out cross-validation
for train_index, test_index in loo.split(X_poly):
X_train, X_test = X_poly[train_index], X_poly[test_index]
y_train, y_test = y[train_index], y[test_index]
# Train the model
model = LinearRegression()
model.fit(X_train, y_train)
# Predict the output
y_pred.append(model.predict(X_test)[0])
y_true.append(y_test[0])
# Calculate accuracy (mean squared error)
mse = mean_squared_error(y_true, y_pred)
accuracy = 1 - mse
print(f"Leave-One-Out Mean Squared Error: {mse}")
print(f"Accuracy: {accuracy}")