Hi Sophy - welcome to UQWorld
I had a look at your code and have a few comments:
- Your Kriging metamodel looks fine and with a global validation error of \varepsilon_{\mathrm{val}}=1.8\% is probably sufficiently accurate to replace your original model, however, to ensure that the metamodel is sufficiently accurate near the point estimator \mathbf{x}_{\mathrm{MAP}} or the posterior mean, I would recommend that you rerun your original model at this point to ensure that it is also sufficiently accurate there;
- The way you define your forward model greatly slows down the MCMC algorithm. You don’t need to reload
Kriging_MetaModel.mat as part of the model definition. Instead, I suggest you load Kriging_MetaModel.mat once and pass it directly to the BayesOpts.Model field;
These are mostly efficiency issues, there are also two bigger issues with your code:
Data
Typically, you would assign the data (i.e., your measurements) directly to the BayesOpts.Data.y field and not mix them with your model definition by letting the model output the difference between the two, but with as little work as possible in your case you would set the data to \mathcal{Y}=0. However, this implicitly assumes, that you only have a single measurement - is this true? Also, setting up your problem this way precludes you from using the default discrepancy options (see below).
Discrepancy options
The discrepancy options specify the properties of the discrepancy between the forward model \mathcal{M} and the data \mathcal{Y} as
y = \mathcal{M}(\mathbf{X}) + \varepsilon
where \varepsilon\sim\mathcal{N}(0,\sigma^2). As you don’t know the proper value for \sigma^2, the Bayesian framework lets you infer the proper value together with the parameters of your forward model.
Note that we deemed your Kriging surrogate model sufficiently accurate and therefore neglect the difference between the actual forward model and the Kriging metamodel!
The simplest way to infer the discrepancy parameters would be not to define a DiscrepancyOptsKnown field and instead use the default options. I suggest you have a quick look at sections 1.2.3 and 2.2.6.2 of the module user manual for details.
In your case of \mathcal{Y}=0, however this will cause an error, because the default prior for \sigma^2 will be \mathcal{U}(0,\mu_{\mathcal{Y}}^2=0). I suggest you define the discrepancy options yourself and use a high enough upper bound for the uniform prior.
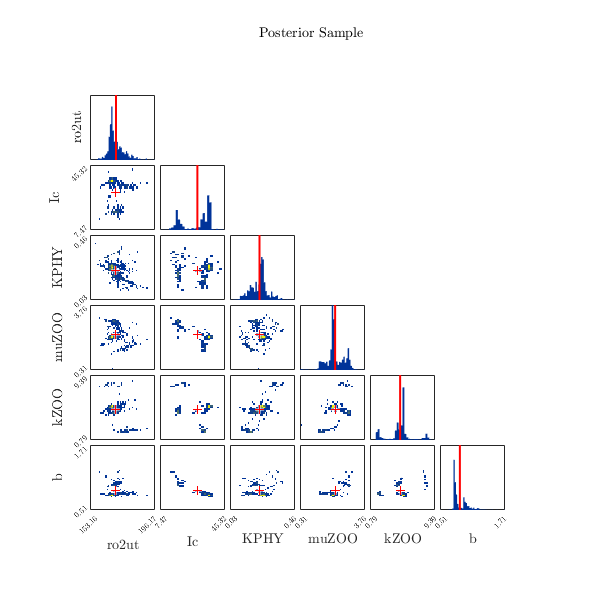
Anyways, I attach an updated version of your file Bayesian_Study_on_Kriging_Model, just let it run and tell me if you are happy with the results.
Bayesian_Study_on_my_Kriging_Model.m (1.8 KB)
P.S.: We just released UQLab version 1.3 - be sure to check it out!